 
SDcols = grep("ID", names(d3), invert = TRUE)] # for all cols other than ID, within each ID value take 1 instead of 0 or highest non-missing instead of missingĭ4 <- d3[, lapply(.SD, max, na.rm = TRUE),īy = ID. 
That helps! OK, using your data I (1) converted both tibbles to data.tables & then changed column1, column2, column10, & column11 to numeric variables using as.numeric() because "N/A" is a character value, whereas NA_real_ is a missing value for a numeric variable. Slice_max(order_by = tibble(across(starts_with("column2"), desc)), n = 1, with_ties = FALSE) This is the code I was using from a very kind reddit user that seems to summarize column 1 and 2 correctly, but turns all of columns 10 and 11 into N/A values for no apparent reason? If anyone can understand why this occurs or has a different method of doing this successfully, i would be very much appreciative. 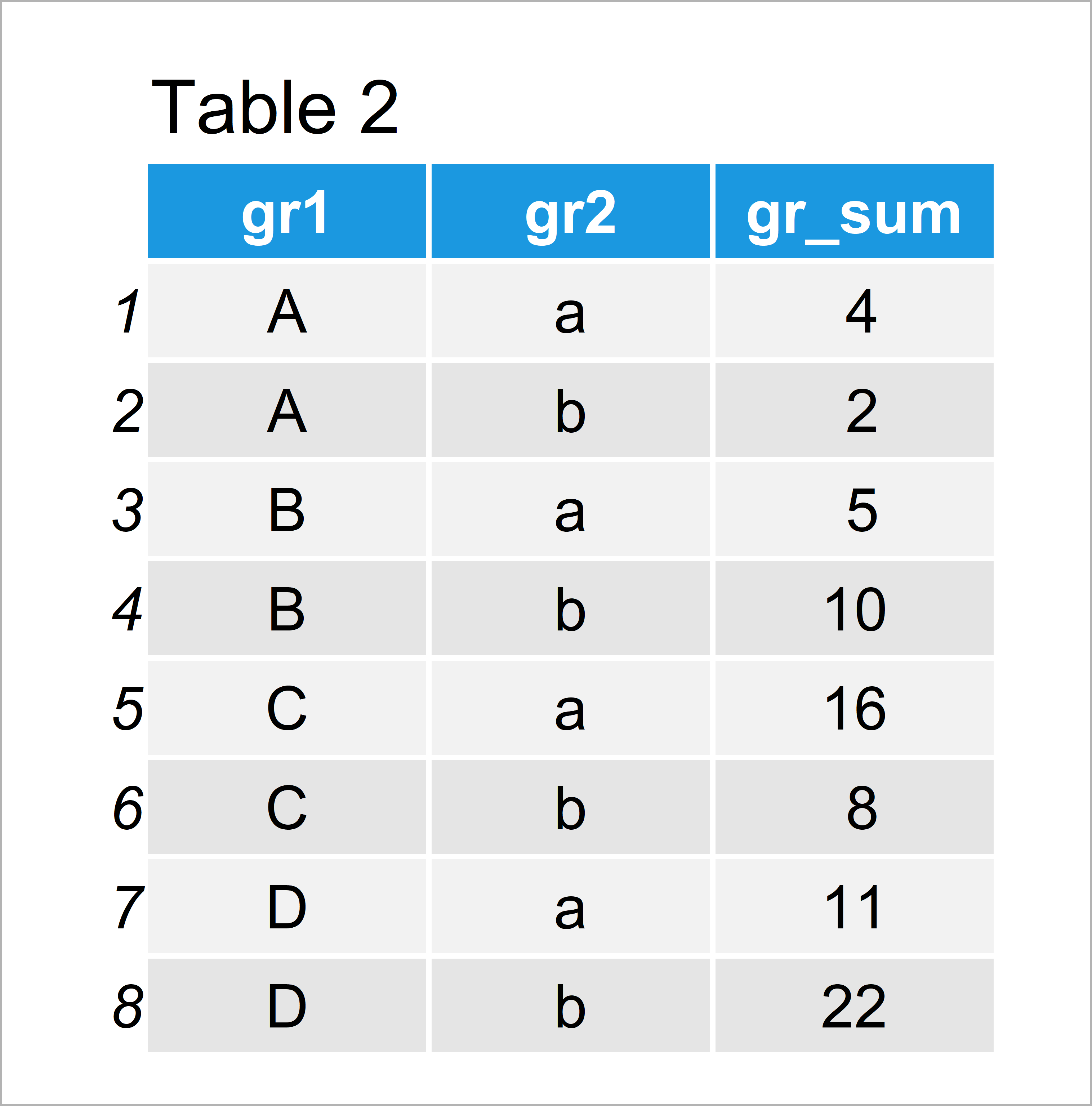
I have tried multiple methods including this code here however, when I do this, for some strange reason, ALL values in columns 10 and 11 (these are columns from df2 originally) turn to N/A. "tbl", "ame"), row.names = c(NA, -28L))įrom df3, my goal is to (see pic below too) summarize all rows per unique ID into one row, with a "1" in any binary column superseding a 0, and using any numerical value present that supersedes a N/A. first, i joined df1 and df2, even including IDs that aren't in both dfs, and made df3 (I inputted NAs for any ID that wasn't present). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
